
A brief overview of the DSS method is available here.
Please ensure that at least 3 sequences have been selected from your alignment before attempting to run DSS. See selecting sequences for further information.
The DSS (Difference of Sums of Squares) analysis method is run by selecting Analysis | Run DSS (Topal-lite) from the menu bar, clicking the Run DSS toolbar button, or pressing Alt-3. This brings up the Difference of Sums of Squares - Confirm Settings dialog.
This dialog provides two groups of parameters that can be customized before starting the DSS run.

Basic settings are available by selecting the Basic tabbed pane.
Depending on the number of sequences that are to be analyzed, there may be up to three analysis methods listed under the Analysis run method drop down box.
Note that no threshold calculations can be performed if either of the latter two options are selected, hence no automatic report generation can be carried out on the results generated from runs using these settings.

The Advanced tabbed pane provides further settings that can be set in order to customize the way in which DSS runs.
Once you are happy with the settings, click Run to start the DSS run. While DSS is running, the DSS Progress dialog will be displayed.
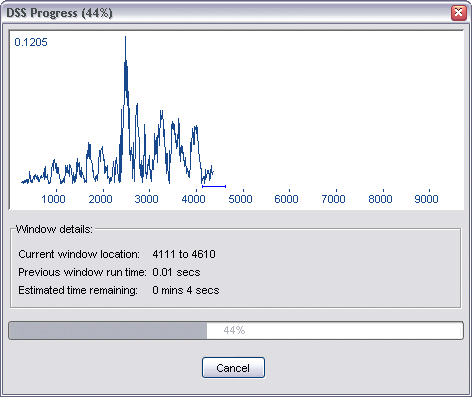
This dialog provides you with a real-time plot of the DSS statistic as the DSS run proceeds, in addition to window information relating to the current window position and the previous window run time.
If a DSS run method has been chosen that allows for threshold calculations, these will take place once the primary DSS run has completed (and the progress bar has reached 100%). The progress bar will then reset to zero and begin to track the threshold calculations. The thresholds are also plotted on the graph once enough processing has occurred to display them, and they will adjust in position as the values become more accurate over time.
Once the DSS run is completed, a dialog will inform you of the total time taken.
The DSS results are then displayed within a DSS Results panel. See graphs for further information on this panel and its results.