TOPALi
TOPALi provides an intuitive graphical interface for working
with existing statistical methods designed for detecting recombination
in DNA sequence alignments. Once recombinant regions have been
identified, further (automated if desired) analysis is then possible,
including the creation of Phylogenetic trees, mosaic structure
predictions, and HTML report generation.
TOPALi has now been superceded by TOPALi v2. Click here for details.
Current version is 0.28, released 21st Nov 2005
(view changelog)

The following is a short list of some of the features and
functionality
TOPALi provides:
- Load DNA multiple alignments in a variety of commonly used formats
- Edit or prune alignments (manually or through phylogeny) to
reduce dataset sizes
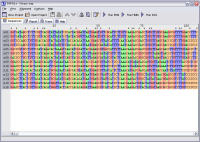
- View alignments with full user-definable colour-coding and scaling
- Run one of three statistical methods to detect recombination on
alignments from 3 to n sequences in size
- PDM (Probabilistic Divergence Measures) and Pruned PDM
- HMM (Hidden Markov Models)
- DSS (Difference of Sums of Squares)
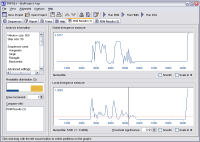
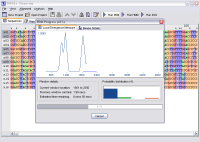
- View results graphically and in real-time as they are generated
(real-time results available with PDM and DSS only)
- Compare and overlay graphs from different runs, and interact with
them to select alignment subsets and partitions
- Partition alignments manually or automatically via a result set's
thresholds, then use those partitions to automatically predict the
alignment's mosaic structure
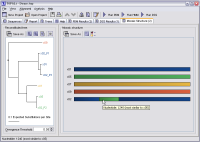
- View the mosaic structure in full-colour, and modify its
parameters to affect how it looks
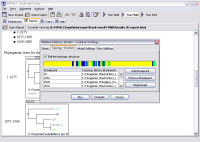
- Create phylogenetic trees for the whole alignment, or just for
sequence or partition subsets
- Export partitions, graphs, trees, and mosaic structure diagrams
in a variety of formats (GIF, SVG, plain text, etc)
- Generate HTML reports that automatically run all of TOPALi's
supported analysis methods for you
You can also view TOPALi's online help
for more information on what is available.
TOPALi is written in (and requires) Java, and should work on
any platform that supports Java 1.4.1 or later. Note however, that its
analysis methods work by calling external C++ programs. These have
currently been compiled and tested on Windows, Unix (Solaris) and Linux
(RH 8 and 9 & Fedora). It may be possible to compile some or all of
them (and hence use those methods within TOPALi) on other
platforms, but we cannot offer any support for this as we have no way
of testing this ourselves.
(TOPALi is released under the GNU General Public License.
See here for further information on
this and the licenses for TOPALi's helper applications).
Selected publications related to this work, include:
- Milne I, Wright F, Rowe G, Marshal DF, Husmeier D and McGuire G
(2004), TOPALi: Software for Automatic Identification of
Recombinant Sequences within DNA Multiple Alignments,
Bioinformatics 20 (11), 1806-1807. View
PDF.
- Husmeier D and McGuire G (2003), Detecting Recombination in
4-Taxa DNA Sequence Alignments with Bayesian Hidden Markov Models and
Markov Chain Monte Carlo, Molecular Biology and Evolution 20 (3),
315-337.
- Husmeier D and Wright F (2001), Probabilistic Divergence
Measures for Detecting Interspecies Recombination, Bioinformatics
17, Suppl. 1, S123-S131.
- McGuire G and Wright F (2000) TOPAL 2.0: Improved Detection
of Mosaic Sequences within Multiple Alignments, Bioinformatics, 16
(2), 130-134.
Please do not hesitate to contact us at topali
(at) bioss.ac.uk if you have any further queries or comments
about this software.





